pacman::p_load(tidyverse, FunnelPlotR, plotly, knitr)package 'FunnelPlotR' successfully unpacked and MD5 sums checked
The downloaded binary packages are in
C:\Users\idrin\AppData\Local\Temp\RtmpeAwTUc\downloaded_packagesKe Ke
April 30, 2024
June 26, 2024
Funnel plot is a specially designed data visualisation for conducting unbiased comparison between outlets, stores or business entities.
Learning Objectives:
plotting funnel plots by using funnelPlotR package,
plotting static funnel plot by using ggplot2 package, and
plotting interactive funnel plot by using both plotly R and ggplot2 packages.
The following R packages will be used:
readr for importing csv into R.
FunnelPlotR for creating funnel plot.
ggplot2 for creating funnel plot manually.
knitr for building static html table.
plotly for creating interactive funnel plot.
Code chunk below will be used to check if these packages have been installed and also will load them into the working R environment.
The COVID-19_DKI_Jakarta will be used. The data was downloaded from Open Data Covid-19 Provinsi DKI Jakarta portal.
For this hands-on exercise, compares the cumulative COVID-19 cases and death by sub-district (i.e. kelurahan) as at 31st July 2021, DKI Jakarta.
The code chunk below imports the data into R and save it into a tibble data frame object called covid19.
FunnelPlotR package uses ggplot to generate funnel plots. It requires a numerator (events of interest), denominator (population to be considered) and group.
The key arguments selected for customisation are:
limit: plot limits (95 or 99).
label_outliers: to label outliers (true or false).
Poisson_limits: to add Poisson limits to the plot.
OD_adjust: to add overdispersed limits to the plot.
xrange and yrange: to specify the range to display for axes, acts like a zoom function.
Other aesthetic components such as graph title, axis labels etc.
The code chunk below plots a funnel plot.
group in this function is different from the scatterplot. Here, it defines the level of the points to be plotted i.e. Sub-district, District or City. If Cityc is chosen, there are only six data points.
By default, data_typeargument is “SR”.
limit: Plot limits, accepted values are: 95 or 99, corresponding to 95% or 99.8% quantiles of the distribution.
A funnel plot object with 267 points of which 0 are outliers. Plot is adjusted for overdispersion.
The code chunk below plots a funnel plot.
data_type argument is used to change from default “SR” to “PR” (i.e. proportions).
x_range and y_range are used to set the range of x-axis and y-axis
A funnel plot object with 267 points of which 7 are outliers. Plot is adjusted for overdispersion.
The code chunk below plots a funnel plot.
label = NA argument is to removed the default label outliers feature.
title argument is used to add plot title.
x_label and y_label arguments are used to add/edit x-axis and y-axis titles.
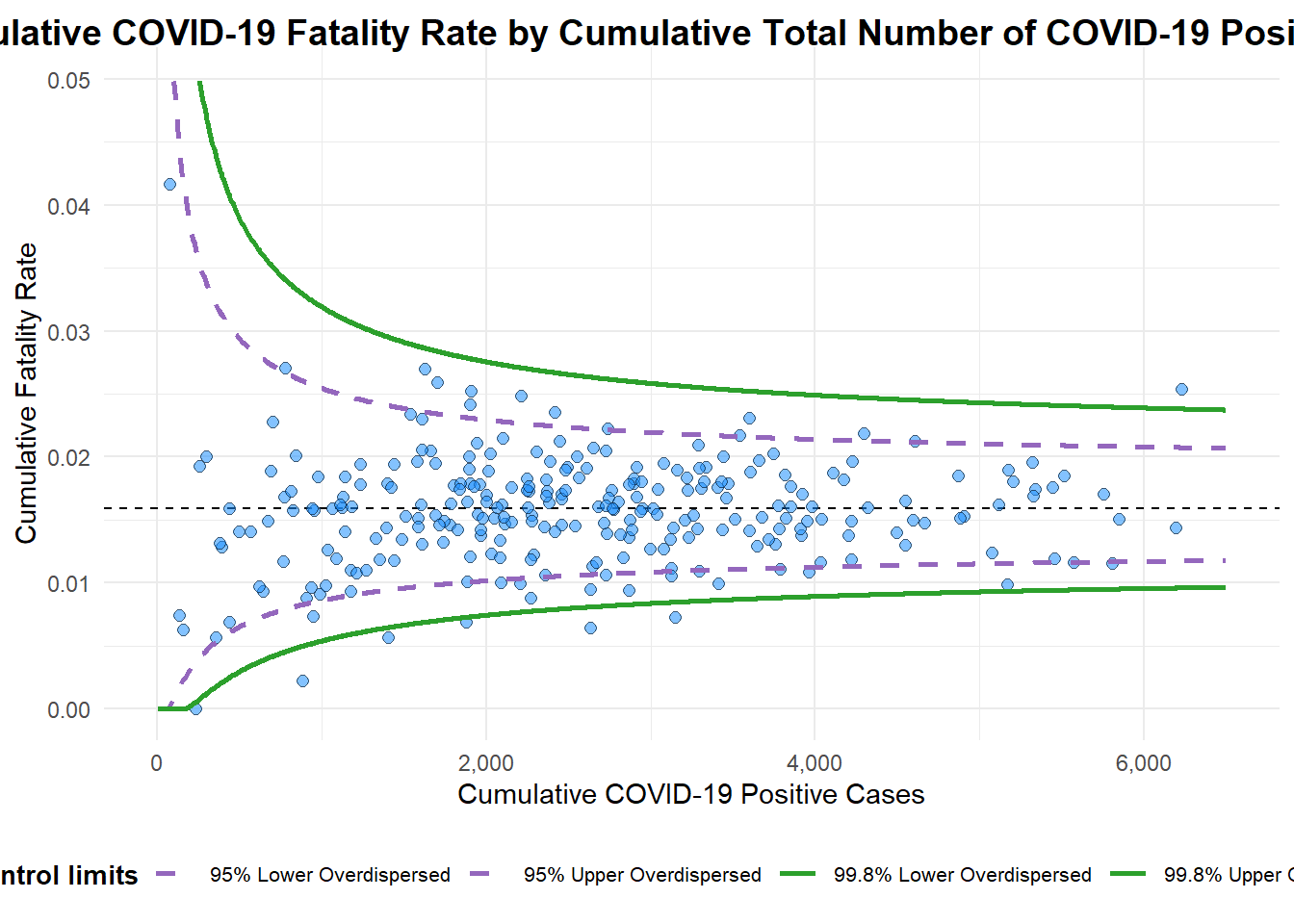
A funnel plot object with 267 points of which 7 are outliers.
Plot is adjusted for overdispersion. funnel_plot(
.data = covid19,
numerator = `Death`,
denominator = `Positive`,
group = `Sub-district`,
data_type = "PR",
x_range = c(0, 6500),
y_range = c(0, 0.05),
label = NA,
title = "Cumulative COVID-19 Fatality Rate by Cumulative Total Number of COVID-19 Positive Cases", #<<
x_label = "Cumulative COVID-19 Positive Cases", #<<
y_label = "Cumulative Fatality Rate" #<<
) A funnel plot object with 267 points of which 7 are outliers. Plot is adjusted for overdispersion.
Build funnel plots using ggplot2
First, derive cumulative death rate and standard error of cumulative death rate.
Next, the fit.mean is computed by using the code chunk below.
The code chunk below is used to compute the lower and upper limits for 95% confidence interval.
number.seq <- seq(1, max(df$Positive), 1)
number.ll95 <- fit.mean - 1.96 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ul95 <- fit.mean + 1.96 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ll999 <- fit.mean - 3.29 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
number.ul999 <- fit.mean + 3.29 * sqrt((fit.mean*(1-fit.mean)) / (number.seq))
dfCI <- data.frame(number.ll95, number.ul95, number.ll999,
number.ul999, number.seq, fit.mean)In the code chunk below, ggplot2 functions are used to plot a static funnel plot.
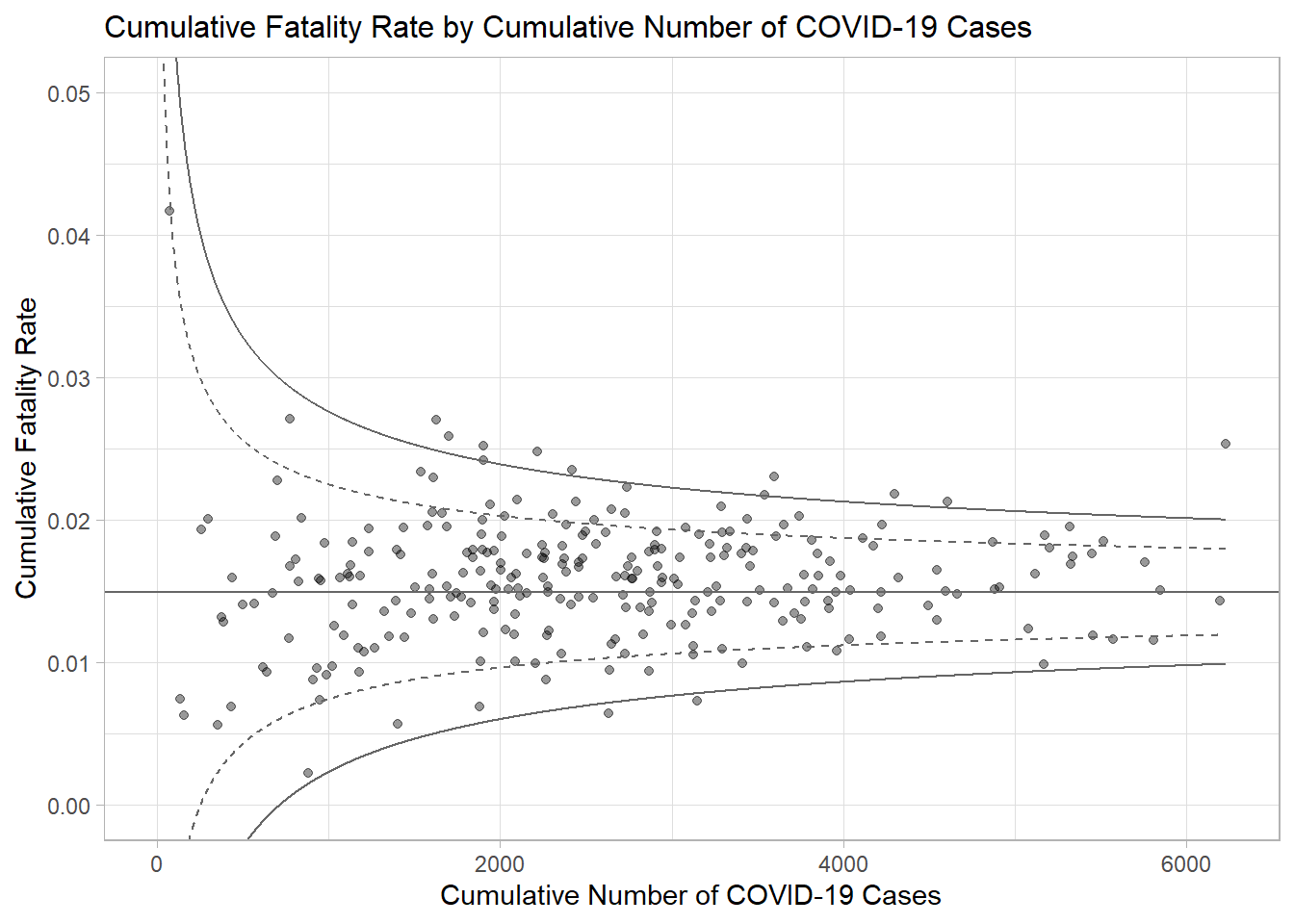
p <- ggplot(df, aes(x = Positive, y = rate)) +
geom_point(aes(label=`Sub-district`),
alpha=0.4) +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ll95),
size = 0.4,
colour = "grey40",
linetype = "dashed") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ul95),
size = 0.4,
colour = "grey40",
linetype = "dashed") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ll999),
size = 0.4,
colour = "grey40") +
geom_line(data = dfCI,
aes(x = number.seq,
y = number.ul999),
size = 0.4,
colour = "grey40") +
geom_hline(data = dfCI,
aes(yintercept = fit.mean),
size = 0.4,
colour = "grey40") +
coord_cartesian(ylim=c(0,0.05)) +
annotate("text", x = 1, y = -0.13, label = "95%", size = 3, colour = "grey40") +
annotate("text", x = 4.5, y = -0.18, label = "99%", size = 3, colour = "grey40") +
ggtitle("Cumulative Fatality Rate by Cumulative Number of COVID-19 Cases") +
xlab("Cumulative Number of COVID-19 Cases") +
ylab("Cumulative Fatality Rate") +
theme_light() +
theme(plot.title = element_text(size=12),
legend.position = c(0.91,0.85),
legend.title = element_text(size=7),
legend.text = element_text(size=7),
legend.background = element_rect(colour = "grey60", linetype = "dotted"),
legend.key.height = unit(0.3, "cm"))
pThe funnel plot created using ggplot2 functions can be made interactive with ggplotly() of plotly r package.